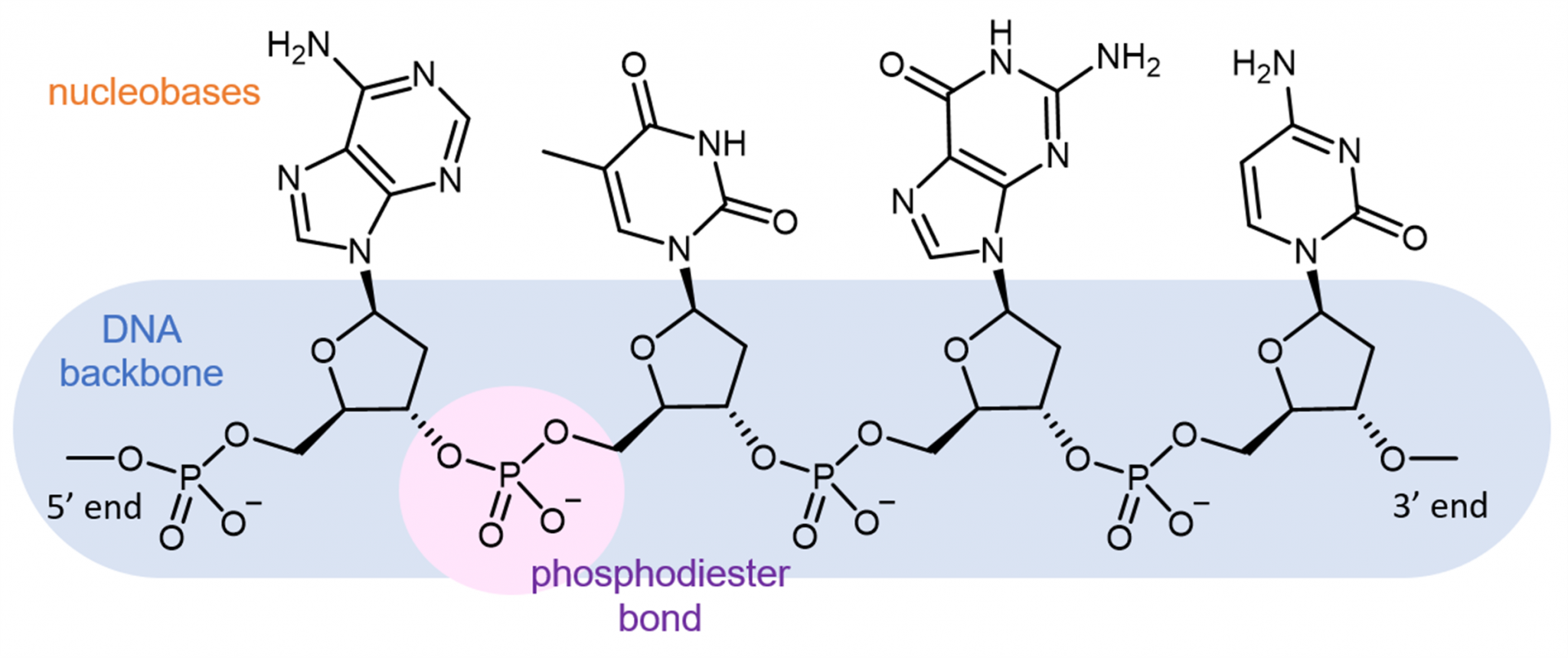
The phosphate group is attached to the 5′ carbon of one nucleotide and the 3′ carbon of the next nucleotide. The carbon atoms of the five-carbon sugar are numbered clockwise from the oxygen as 1′, 2′, 3′, 4′, and 5′ (1′ is read as “one prime”). The sugar–phosphate groups line up in a “backbone” for each single strand of DNA, and the nucleotide bases stick out from this backbone. The phosphate group of one nucleotide bonds covalently with the sugar molecule of the next nucleotide, and so on, forming a long polymer of nucleotide monomers. Figure 9.3 (a) Each DNA nucleotide is made up of a sugar, a phosphate group, and a base.įigure 9.3 (b) Cytosine and thymine are pyrimidines. The nucleotide is named according to the nitrogenous base it contains. Adenine (A) and guanine (G) are double-ringed purines, and cytosine (C) and thymine (T) are smaller, single-ringed pyrimidines. There are four types of nitrogenous bases in DNA. The building blocks of DNA are nucleotides, which are made up of three parts: a deoxyribose (5-carbon sugar), a phosphate group, and a nitrogenous base ( Figure 9.3). Now let’s consider the structure of the two types of nucleic acids, deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). (credit a: modification of work by Marjorie McCarty b: modification of work by NIH) Scientist Rosalind Franklin discovered (b) the X-ray diffraction pattern of DNA, which helped to elucidate its double helix structure. Figure 9.2 Pioneering scientists (a) James Watson and Francis Crick are pictured here with American geneticist Maclyn McCarty. In 1962, James Watson, Francis Crick, and Maurice Wilkins were awarded the Nobel Prize in Medicine for their work in determining the structure of DNA. This meant they were always paired in some way. Chargaff had shown that of the four kinds of monomers (nucleotides) present in a DNA molecule, two types were always present in equal amounts and the remaining two types were also always present in equal amounts. Watson and Crick also had key pieces of information available from other researchers such as Chargaff’s rules. Watson and Crick were able to piece together the puzzle of the DNA molecule using Franklin’s data ( Figure 9.2). In Wilkins’ lab, researcher Rosalind Franklin was using X-ray crystallography to understand the structure of DNA. The patterns give important information about the structure of the molecule of interest. X-ray crystallography is a method for investigating molecular structure by observing the patterns formed by X-rays shot through a crystal of the substance. Pauling had discovered the secondary structure of proteins using X-ray crystallography. Other scientists, such as Linus Pauling and Maurice Wilkins, were also actively exploring this field. In the 1950s, Francis Crick and James Watson worked together at the University of Cambridge, England, to determine the structure of DNA. Describe how eukaryotic and prokaryotic DNA is arranged in the cell.School of Chemistry and Biochemistry, Georgia Institute of Technology, Atlanta, Georgia 30332-0400, USA.By the end of this section, you will be able to: 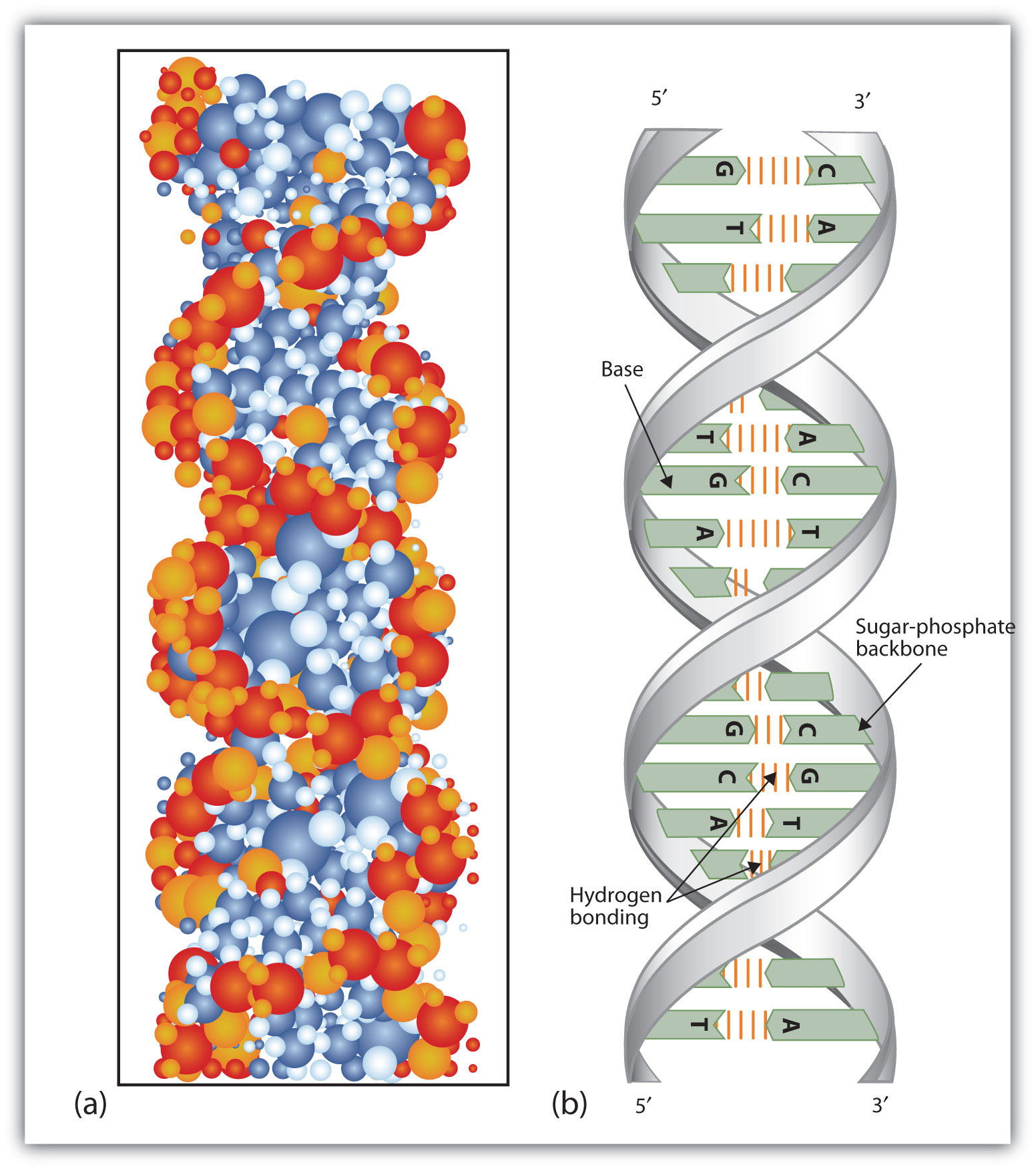
This bending may be related to the direct coordination of a sodium cation by a DNA base, with unprecedented inner-shell (direct) coordination of penta-hydrated sodium at the O6 atom of a guanine. Electrostatic forces appear to induce modest DNA bending into the major groove. TriplatinNC extends along the phosphate backbone, in a mode of binding we call "Backbone Tracking" and spans the minor groove in a mode of binding we call "Groove Spanning". The high repetition and geometric regularity of the motif suggests that this type of Pt(II) center can be developed as a modular nucleic acid binding device with general utility. The interaction appears to prefer O2P over O1P atoms (frequency of interaction is O2P > O1P, base and sugar oxygens > N). The geometry is conserved among the 8 observed phosphate clamps in this structure. The three square-planar tetra-am(m)ine Pt(II) coordination units form bidentate N.O.N complexes with OP atoms, in a motif we call the Phosphate Clamp. Instead, it binds to phosphate oxygen atoms and thus associates with the backbone. TriplatinNC does not intercalate nor does it bind in either groove. TriplatinNC is a multifunctional DNA ligand, with three cationic Pt(II) centers, and directional hydrogen bonding functionalities, linked by flexible hydrophobic segments, but without the potential for covalent interaction. We describe a 1.2 A X-ray structure of a double-stranded B-DNA dodecamer (the Dickerson Dodecamer, DDD, 2) associated with a cytotoxic platinum(II) complex, (TriplatinNC). Diversity, Equity, Inclusion, and Access.Citation, Usage, Privacy Policies, Logo.Biologically Interesting Molecule Reference Dictionary (BIRD).
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
